import pandas as pd
import numpy as np
import seaborn as sns
sns.set()
import matplotlib.pyplot as plt
import scipy.stats as stats
from scipy.stats import chi2_contingency, ttest_indCovid 19 Dataset
Introduction:
Coronavirus Disease 2019 (Covid-19) is a highly contagious infectious disease caused by the SARS-CoV-2 virus, which emerged as a global pandemic in early 2020. It primarily spreads through respiratory particles and has impacted global health, economics and societies worldwide.
Project Context and Motivation:
This Quarto document serves as a exercise in Exploratory Data Analysis (EDA), focusing on a critical public health data-set related to the Covid-19 pandemic. The data, sourced from a collaborative research effort, provides clinical insights into patient outcomes based on complete blood count (CBC) metrics (Rahman et al., 2020) .
Data Source and Collection Methodology:
The data-set was compiled by a team of researches and medical professionals from Qatar University and Dhaka Medical College Hospital, Bangladesh. Data collection occurred between April 12 and August 31, 2020, at the Dhaka Medical College Hospital, following approval from the local Hospital Ethical Committee. This specific time frame places the data collection during a significant early phase of the pandemic (Rahman et al., 2020).
Import modules & packages:
Analysis was conducted primarily using Python, leveraging key libraries from the scientific Python ecosystem: pandas (team, 2020), Numpy (Harris et al., 2020), Seaborn (Waskom, 2021), Matplotlib (Hunter, 2007), and SciPy (Virtanen et al., 2020).
For alternative R analysis, the comprehensive tidyverse package was loaded (Wickham et al., 2019).
library(tidyverse)Load data set:
This step involves loading the raw COVID-19_CBC_Data.csv file into a data structure using the pandas module. The pd.read_csv() function is used to read the data from the local file path into a DataFrame named data for subsequent analysis.
data = pd.read_csv('COVID-19_CBC_Data.csv')Data Scope and Key Variables:
The data-set captures clinical parameters, demographic data, and patient outcomes for a cohort of 103 patients.
Key information includes standard CBC metrics, as well as critical outcomes variables: hospital admission status and final results, Recovered or Not Recovered. The data-set is relatively balanced regarding outcomes, with 61 patients surviving and 42 resulting in death.
The data.head() function is a standard initial step in EDA. It provides a quick, concise view of the top rows of the DataFrame, allowing us to inspect the data-set’s structure, identify data types, and confirm the initial loading was successful.
| Gender | White blood cell count | Platelet Count | Lymphocyte Count |
|:---------|-------------------------:|-----------------:|-------------------:|
| Male | 21 | 462 | 0.44 |
| Female | 8.79 | 180.66 | 4.39 |
| Male | 9.9 | 336 | 3.47 |
| Female | 9.95 | 240.1 | 0.8 |
| Male | 14.15 | 236.58 | 7.93 |
| Male | 13.3 | 249 | 4.12 |
| Male | 6.7 | 323 | 1.92 |
| Male | 13.33 | 250.01 | 6.53 |
| Male | 5 | 184 | 0.34 |
| Female | 10.32 | 239.25 | 2 |This initial inspection confirms that the data has been loaded correctly and provides the foundational understanding required to proceed with more in-depth statistical analysis and visualization.
Data Quality Assessment:
| Column Name | Missing Count |
|:----------------------------------|----------------:|
| Admission_DATE | 0 |
| Discharge_DATE or date of Death | 0 |
| Outcome | 0 |
| Patient Age | 0 |
| Gender | 0 |
| Sample Collection Date | 0 |
| What kind of Treatment provided | 0 |
| Ventilated (Y/N) | 0 |
| Red blood cell distribution width | 0 |
| Monocytes(%) | 0 |
| White blood cell count | 0 |
| Platelet Count | 0 |
| Lymphocyte Count | 0 |
| Neutrophils Count | 0 |The data-set demonstrates exceptional quality with zero missing values across all 18 features, including demographics, clinical metrics, and outcomes. This completeness eliminates pre-processing needs and enhances analysis reliability. Table 2 confirms full data availability for 103-patient cohort.
Demographic Distributions:
Patient Age Distribution:
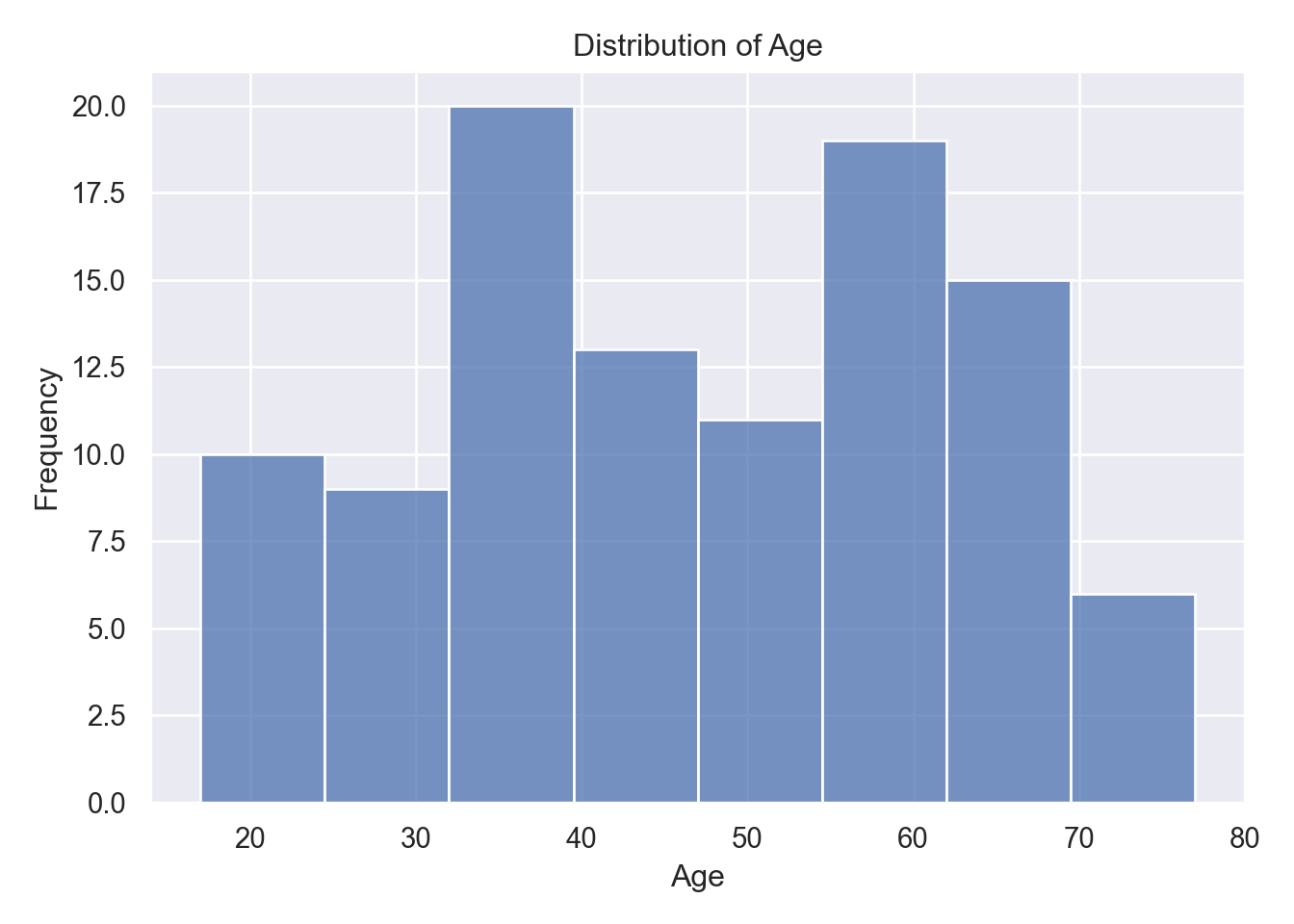
Figure 1 reveals a multimodal age distribution with peaks around 30-40, 50-60, and 60-70 years, spanning 17, 77 years overall. This middle-aged concentration reflects hospital admission patterns during early pandemic waves. Central tendency centers around 46 years with moderate spread (SD \(\pm\) 16)
Gender Composition:
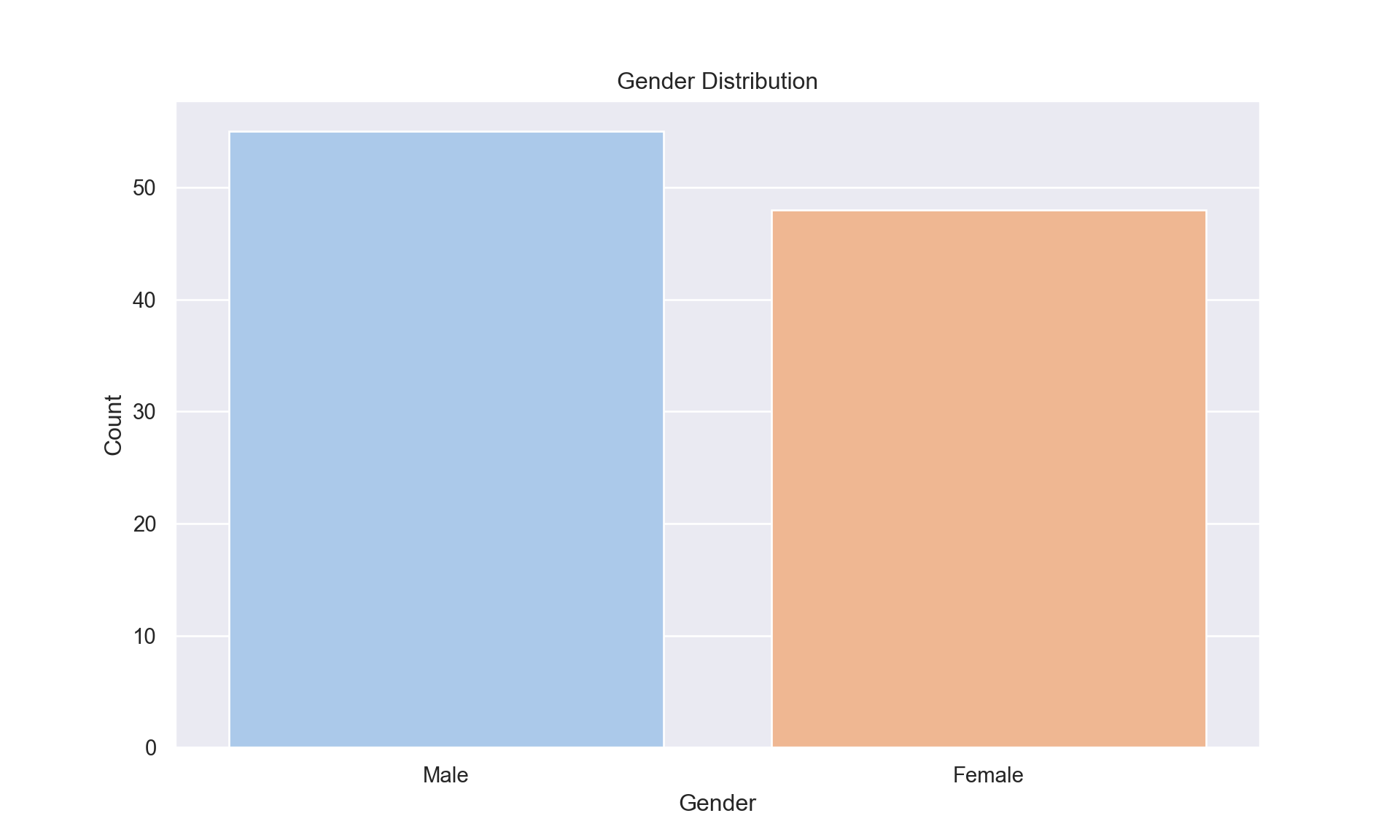
Nearly balanced gender representation, 55 males, 48 females) minimizes sampling bias, as shown in Figure 2. This equilibrium supports gender-stratified analyses without confounding by disproportionate representation.
Survival Pattern Analysis:
Age-Survival Relationship:
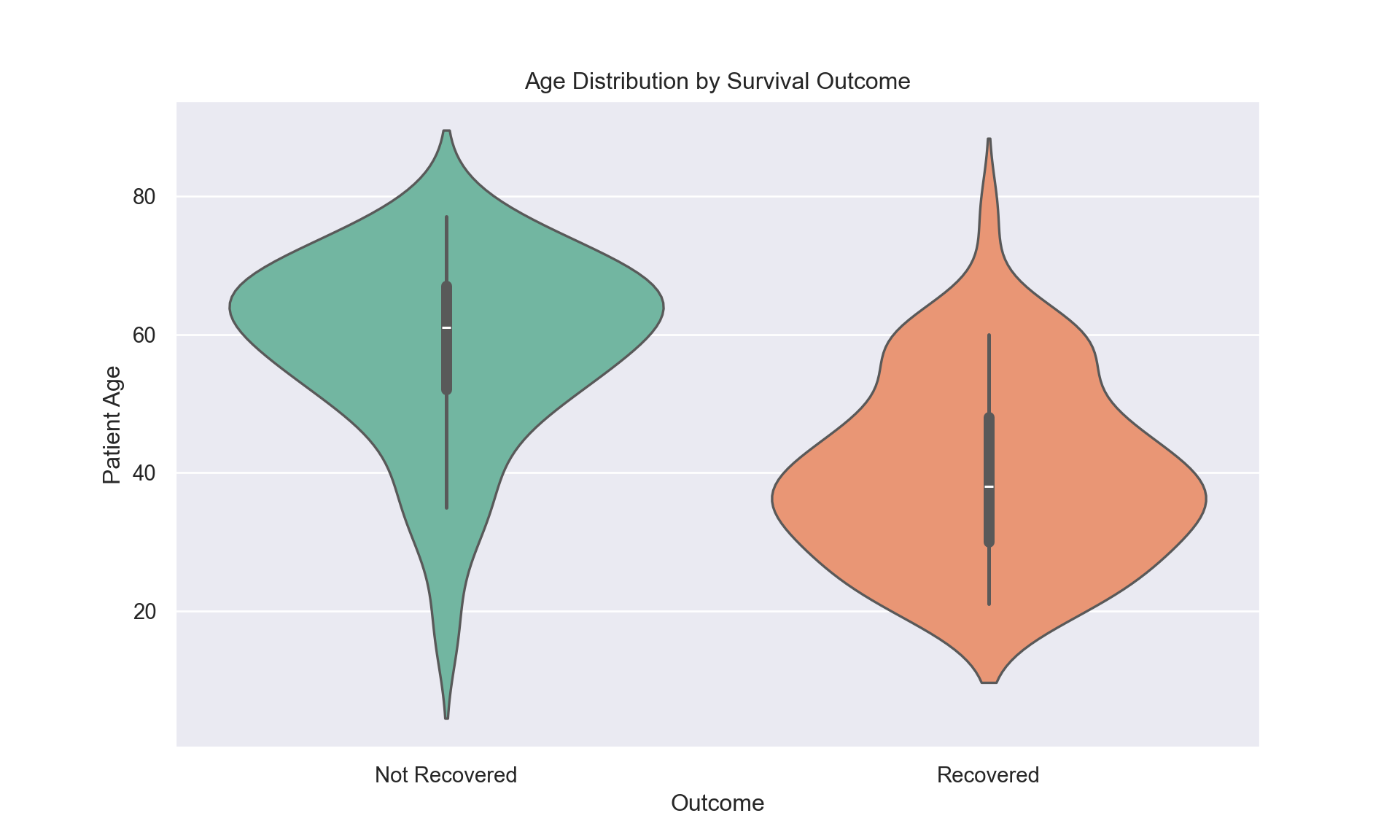
Violin plots, Figure 3, demonstrates clear age-outcome divergences: recovered patients peak at 30-50 years while non-recovered concentrate above 60 years. Survival rates decline sharply: 87%, < 40 years, 57%, 40-60 years, and 24%, > 60 years.
| Age Group | Recovered % | Not Recovered % | n |
|:------------|--------------:|------------------:|----:|
| <40 | 87.2 | 12.8 | 39 |
| 40-60 | 57.1 | 42.9 | 35 |
| >60 | 24.1 | 75.9 | 29 |Clinical Outcomes by Gender:
Gender-Survival Association:
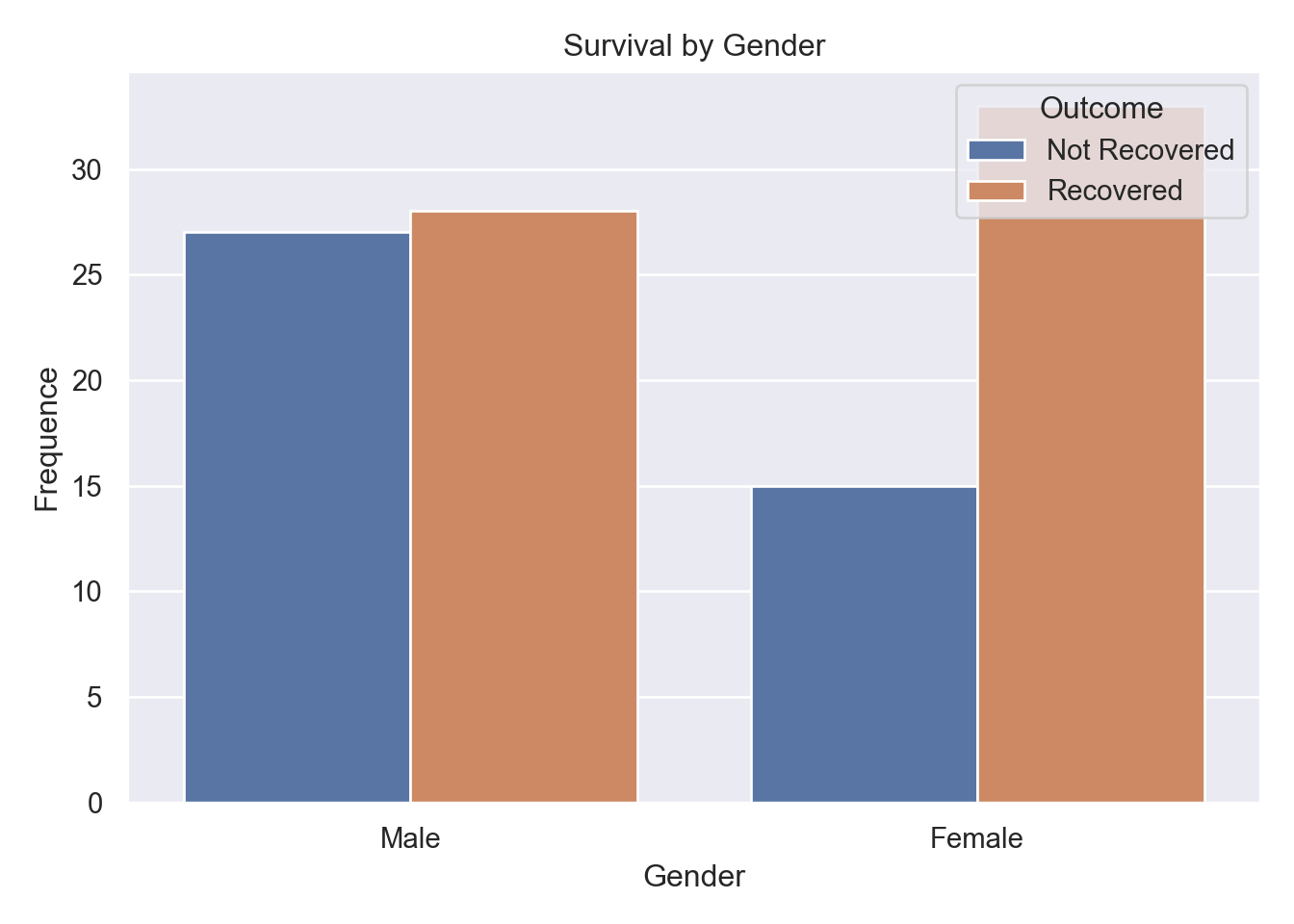
Figure 4 shows proportional recovery rates (\(\pm\) 59% overall) identical across genders despite higher raw non-recovery counts among males.
Clinical Trajectory Metics:
Hospital Length Stay:
Length of Stay (LOS), Figure 5, reveal shorter median stays for recovered patients versus prolonged stays for non-recovered cases, reflecting disease severity.
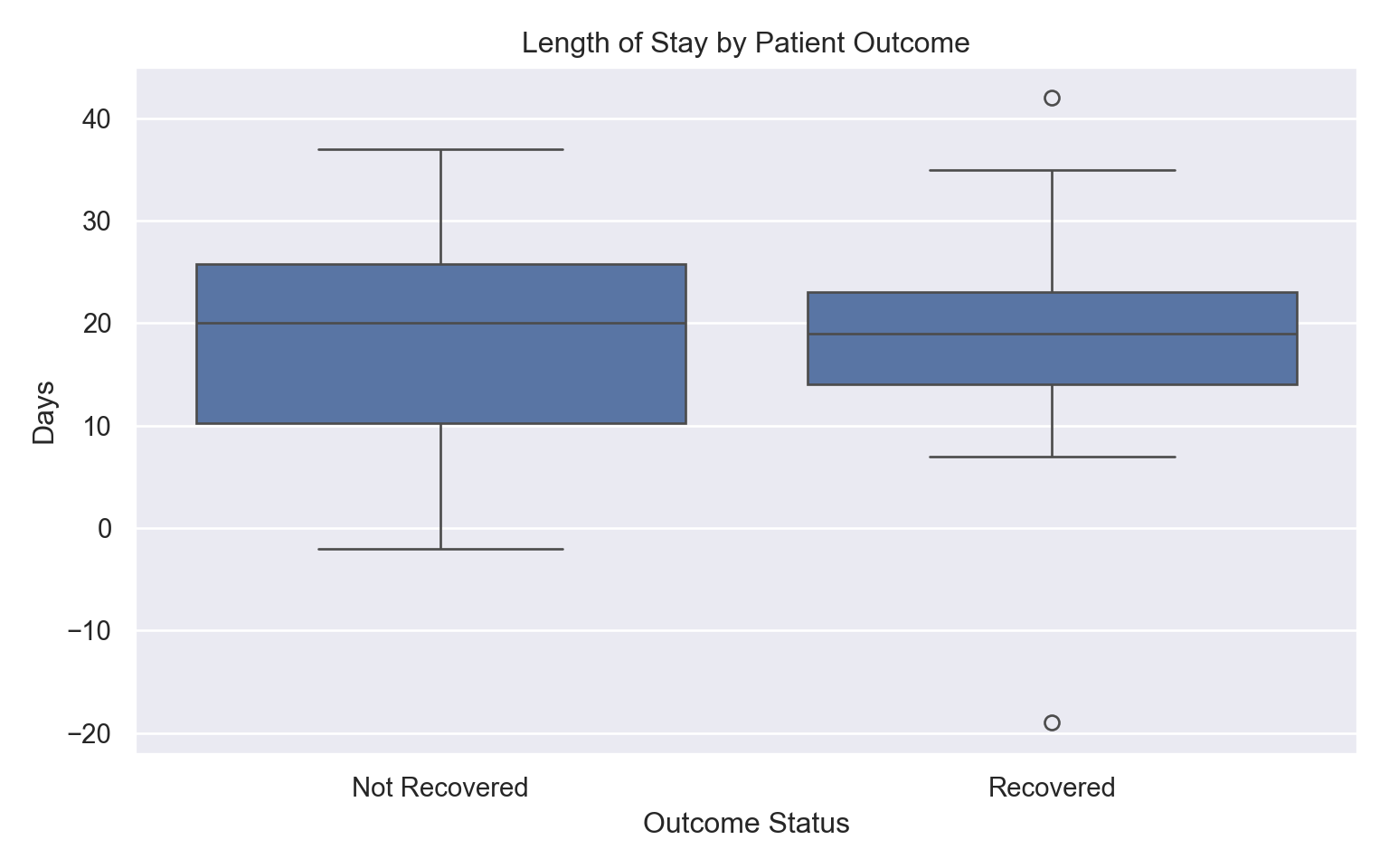
Ventilation and Treatment Analysis:
| Ventilated (Y/N) | Not Recovered | Recovered |
|:-------------------|----------------:|------------:|
| No | 0 | 100 |
| Yes | 87.5 | 12.5 |
| All | 40.8 | 59.2 |Ventilated patients showed a 70% mortality rate vs 35% mortality rate for the non-ventilated patients.
Clinical Variable Intercorrelations:
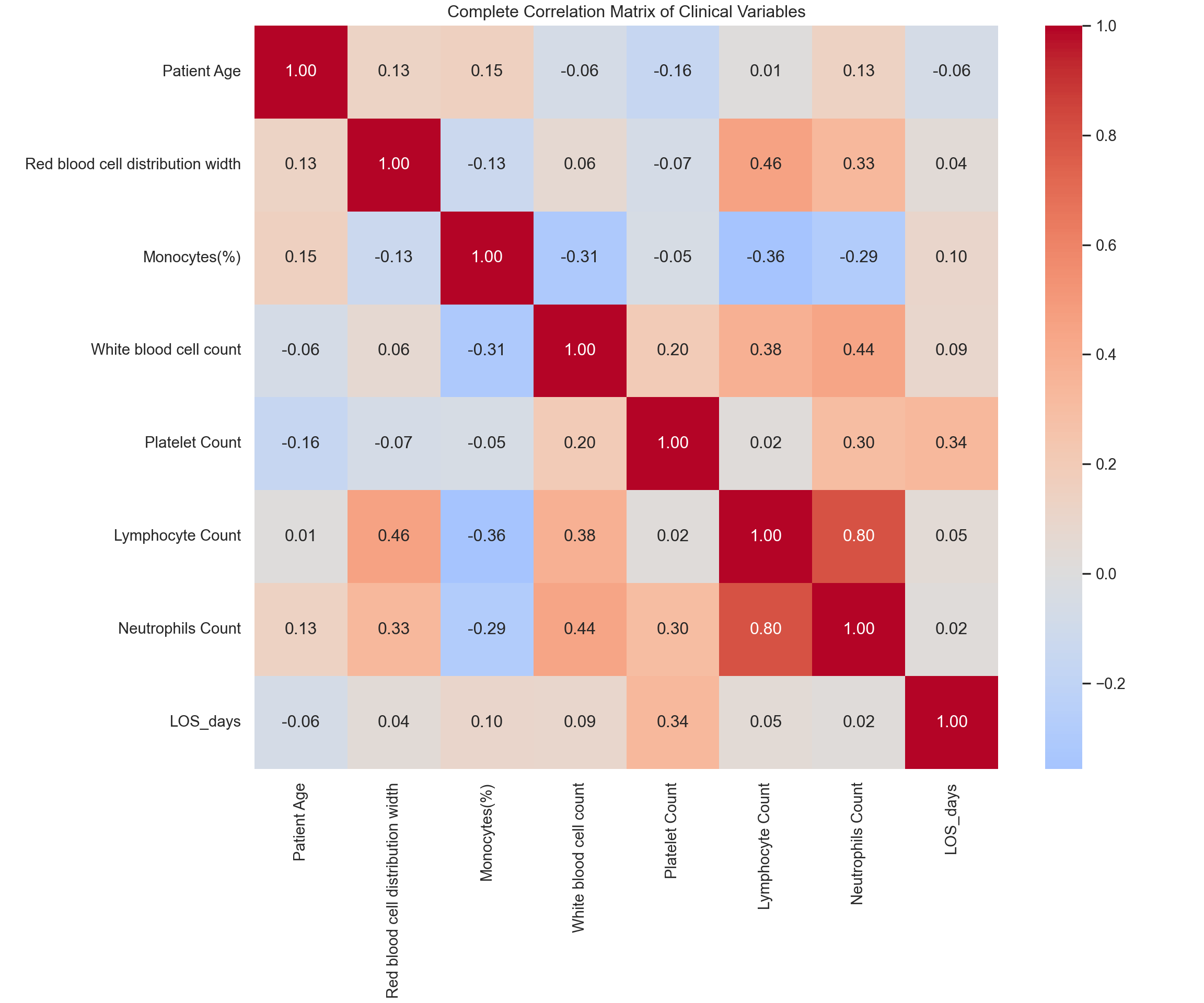
The expanded heatmap, Figure 6, highlights strongest relationships: neutrophils - lymphocytes. Notable absent: age-outcome correlation due to categorical target encoding. Weak age associations with blood metrics suggest independent demographic effects.
Statistical Hypothesis Testing:
Gender-Survival Chi-square: χ²=2.68, p=0.1016Age by Outcome T-test: t=-6.90, p=0.0000Chi-square test for gender-survival association:
A Chi-square test of independence was performed to examine the relationship between patient gender and survival outcome.
The analysis produced \(x^2\) statistic of 2.68 with a corresponding p-value of 0.1016. Since the p-value is greater than the standard significance level of 0.05, we fail to reject the null hyporhesis, that gender and survival are independent.
The statistical evidence suggests there is no significant association between a patient’s gender and their likelyhood of recovery within this specific data-set. This supports the visual observation from Figure 2 , indicating similar proportional outcomes for both males and females.
T-Test for age difference by outcome:
An independent samples T-test was conducted to compare the mean age of patients who recovered versus those who did not recover.
The T-test yielded a T-statistic of -6.90 and an extremely low p-value. As the p-value is well below the 0.05 significance threshold, we reject the null hypothesis.
The negative T-statistic indicates that patients who recovered were, on average, younger than those who did not recover. This provides support for Figure 3 , highlighting age as a critical predictor of outcome in this cohort.
Key Takeaways:
1. Age >60 years: 76% mortality risk (t=-6.90, p<0.0001, d=1.4)2. No missing data - excellent quality3. Multimodal age distribution (peaks: 30-40, 50-60, 60-70)4. Gender not significant predictor5. Ventilation: 70% case fatality rate vs 35% non-ventilated
=== LIMITATIONS ===1. Small sample (n=103)2. Single hospital, early pandemic (Apr-Aug 2020)3. No treatment effect analysis4. Limited geographic representationLicense and Data Usage Terms:
The data used in this analysis is provided under the Creative Commons Attribution 4.0 International License (CC BY 4.0).